Assemblies
Assembly Name:
Protein sprouty homolog 4
Multimeric state:
monomeric
Accessible surface area:
1137.12 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-189877
Assembly Name:
E3 ubiquitin-protein ligase CBL
Multimeric state:
monomeric
Accessible surface area:
14790.8 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-149756
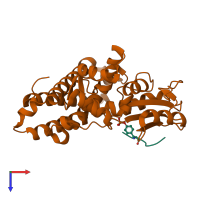
Multimeric state:
hetero dimer
Accessible surface area:
14867.47 Å2
Buried surface area:
1060.46 Å2
Dissociation area:
530.23
Å2
Dissociation energy (ΔGdiss):
6.02
kcal/mol
Dissociation entropy (TΔSdiss):
6.97
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-149768
Macromolecules
Chain: A
Length: 13 amino acids
Theoretical weight: 1.58 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Length: 13 amino acids
Theoretical weight: 1.58 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 Q9C004 (Residues: 46-58; Coverage: 4%)
Q9C004 (Residues: 46-58; Coverage: 4%)
Chain: B
Length: 329 amino acids
Theoretical weight: 38.19 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
SCOP:
Length: 329 amino acids
Theoretical weight: 38.19 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P22681 (Residues: 23-351; Coverage: 36%)
P22681 (Residues: 23-351; Coverage: 36%)
Pfam:
- CBL proto-oncogene N-terminal domain 1
- CBL proto-oncogene N-terminus, EF hand-like domain
- CBL proto-oncogene N-terminus, SH2-like domain
- Adaptor protein Cbl
- Adaptor protein Cbl, PTB domain
- Adaptor protein Cbl, N-terminal domain superfamily
- Adaptor protein Cbl, N-terminal helical
- EF-hand domain pair
- Adaptor protein Cbl, EF hand-like
- Adaptor protein Cbl, SH2-like domain
- SH2 domain superfamily
SCOP: