Function and Biology Details
Reaction catalysed:
Endonucleolytic cleavage of DNA to give specific double-stranded fragments with terminal 5'-phosphates
Biochemical function:
Biological process:
Cellular component:
- not assigned
Structure analysis Details
Assemblies composition:
Assembly name:
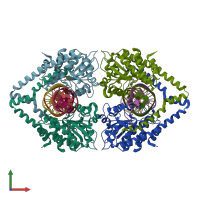
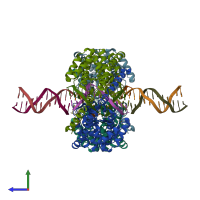
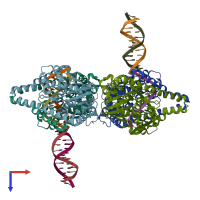
Type II restriction enzyme HindIII and DNA (preferred)
PDBe Complex ID:
PDB-CPX-117098 (preferred)
Entry contents:
1 distinct polypeptide molecule
1 distinct DNA molecule
1 distinct DNA molecule
Macromolecules (2 distinct):