EMD-5130
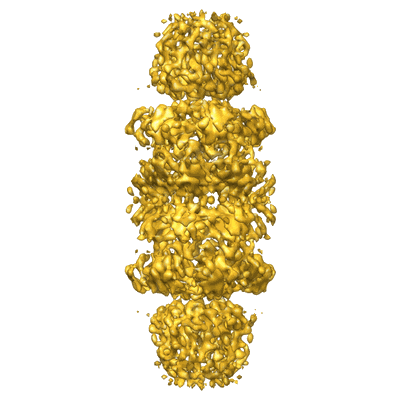
3D cryo-EM Structure of Archaeal 20S Proteasome in Complex with the C-terminus of PAN
EMD-5130
Single-particle7.5 Å
 Deposition: 18/08/2009
Deposition: 18/08/2009Map released: 21/12/2009
Last modified: 23/09/2011
Concentration: 0.3
mg/mL
Buffer
pH: 8.5
Details: 50 mM Tris pH 8.5, 1mM DTT, 10 mM MgCl2, 5% Glycerol
Details: 50 mM Tris pH 8.5, 1mM DTT, 10 mM MgCl2, 5% Glycerol
Grid
Details: 400 mesh Cu grids
Microscope: FEI TECNAI F20
Illumination mode: FLOOD BEAM
Imaging mode: BRIGHT FIELD
Electron source: FIELD EMISSION GUN
Acceleration voltage: 200 kV
Nominal CS: 2.0 mm
Nominal defocus: 1.0 µm - 5.0 µm
Nominal magnification: 62000.0
Specimen holder model: GATAN LIQUID NITROGEN
Specimen holder details: Gatan CT-3500
Minimum tilt angle: 0
Maximum tilt angle: 0
Details: Data was collected using Leginon
Illumination mode: FLOOD BEAM
Imaging mode: BRIGHT FIELD
Electron source: FIELD EMISSION GUN
Acceleration voltage: 200 kV
Nominal CS: 2.0 mm
Nominal defocus: 1.0 µm - 5.0 µm
Nominal magnification: 62000.0
Specimen holder model: GATAN LIQUID NITROGEN
Specimen holder details: Gatan CT-3500
Minimum tilt angle: 0
Maximum tilt angle: 0
Details: Data was collected using Leginon
Temperature
Minimum: 90
K
Average: 90 K
Maximum: 90 K
Average: 90 K
Maximum: 90 K
Image Recording
[1]
Detector category:
CCD
Detector model: GENERIC GATAN (4k x 4k)
Average electron dose per image: 20 e/Å2
Details: Images were recorded on CCD camera
Detector model: GENERIC GATAN (4k x 4k)
Average electron dose per image: 20 e/Å2
Details: Images were recorded on CCD camera
Details: The particles were manually selected
Final
reconstruction
Resolution: 7.5
Å
(
BY AUTHOR)
Resolution method: FSC 0.143 CUT-OFF
Number of images used: 13020
Algorithm: OTHER
Details: D7 symmetry was applied
Resolution method: FSC 0.143 CUT-OFF
Number of images used: 13020
Algorithm: OTHER
Details: D7 symmetry was applied
⌯ Applied Symmetry
Point group:
D7
Software
[1]
| Name | Version | Details |
|---|---|---|
| FREALIGN | - | - |
CTF correction
Details:Each particle
Format: CCP4
Data type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Annotation details: This is the 3D volume of T.acidophilum 20S proteasome in complex with a hybrid activator of PA26 and PAN
Details: ::::EMDATABANK.org::::EMD-5130::::
Data type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Annotation details: This is the 3D volume of T.acidophilum 20S proteasome in complex with a hybrid activator of PA26 and PAN
Details: ::::EMDATABANK.org::::EMD-5130::::
⬡ Geometry
| X | Y | Z | |
|---|---|---|---|
| Dimensions | 116 | 116 | 196 |
| Origin | -58 | -58 | -98 |
| Spacing | 116 | 116 | 196 |
| Voxel size | 1.79103 Å | 1.79103 Å | 1.79102 Å |
Contour list
| Primary | Level | Source |
|---|---|---|
| True | 1.3 | AUTHOR |
