Function and Biology Details
Biochemical function:
- not assigned
Biological process:
- not assigned
Cellular component:
Sequence domains:
Structure analysis Details
Assembly composition:
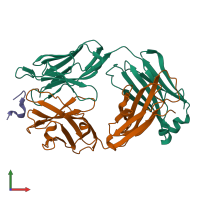
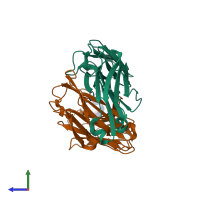
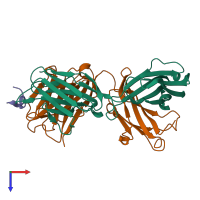
hetero trimer (preferred)
PDBe Complex ID:
PDB-CPX-208679 (preferred)
Entry contents:
3 distinct polypeptide molecules
Macromolecules (3 distinct):
Ligands and Environments
No bound ligands
No modified residues
Experiments and Validation Details
wwPDB Validation report is not available for this entry.
Spacegroup:
C2221
Expression system: Not provided