Function and Biology Details
Reaction catalysed:
(1a) S-adenosyl-L-methionine + 2-((3S)-3-carboxy-3-aminopropyl)-L-histidine-[translation elongation factor 2] = S-adenosyl-L-homocysteine + 2-((3S)-3-carboxy-3-(methylamino)propyl)-L-histidine-[translation elongation factor 2]
Biochemical function:
Biological process:
Cellular component:
- not assigned
Structure analysis Details
Assemblies composition:
Assembly name:
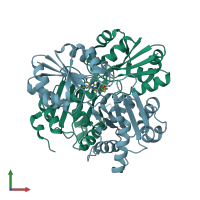
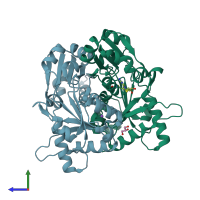
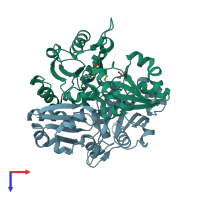
Diphthine synthase (preferred)
PDBe Complex ID:
PDB-CPX-129728 (preferred)
Entry contents:
1 distinct polypeptide molecule
Macromolecule: